FMRP Rabbit mAb [n3Tx]Cat NO.: A43268
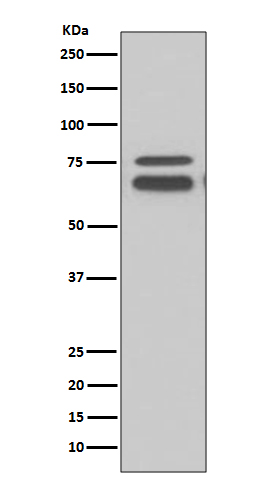
Western blot(SDS PAGE) analysis of extracts from K562 cell lysate.Using FMRP Rabbit mAb [n3Tx]at dilution of 1:1000 incubated at 4℃ over night.
Product information
Protein names :FMR1; Fmr1 gene; FMRP; Fragile X mental retardation 1; Fragile X mental retardation 1 protein; FRAXA;POF; POF1; Protein FMR-1; Protein FMR1;
UniProtID :Q06787
MASS(da) :71,174
MW(kDa) :71kDa
Form :Liquid
Purification :Affinity-chromatography
Host :Rabbit
Isotype : IgG
sensitivity :Endogenous
Reactivity :Human,Mouse,Rat
- ApplicationDilution
- 免疫印迹(WB)1:1000-2000
- 免疫组化(IHC)1:100
- 免疫荧光(ICC/IF)1:100
- The optimal dilutions should be determined by the end user
Specificity :Antibody is produced by immunizing animals with A synthesized peptide derived from human FMRP
Storage :Antibody store in 10 mM PBS, 0.5mg/ml BSA, 50% glycerol. Shipped at 4°C. Store at-20°C or -80°C. Products are valid for one natural year of receipt.Avoid repeated freeze / thaw cycles.
WB Positive detected :K562 cell lysate.
Function : Multifunctional polyribosome-associated RNA-binding protein that plays a central role in neuronal development and synaptic plasticity through the regulation of alternative mRNA splicing, mRNA stability, mRNA dendritic transport and postsynaptic local protein synthesis of a subset of mRNAs (PubMed:16631377, PubMed:18653529, PubMed:19166269, PubMed:23235829, PubMed:25464849). Plays a role in the alternative splicing of its own mRNA (PubMed:18653529). Plays a role in mRNA nuclear export (By similarity). Together with export factor NXF2, is involved in the regulation of the NXF1 mRNA stability in neurons (By similarity). Stabilizes the scaffolding postsynaptic density protein DLG4/PSD-95 and the myelin basic protein (MBP) mRNAs in hippocampal neurons and glial cells, respectively,this stabilization is further increased in response to metabotropic glutamate receptor (mGluR) stimulation (By similarity). Plays a role in selective delivery of a subset of dendritic mRNAs to synaptic sites in response to mGluR activation in a kinesin-dependent manner (By similarity). Plays a role as a repressor of mRNA translation during the transport of dendritic mRNAs to postsynaptic dendritic spines (PubMed:11532944, PubMed:11157796, PubMed:12594214, PubMed:23235829). Component of the CYFIP1-EIF4E-FMR1 complex which blocks cap-dependent mRNA translation initiation (By similarity). Represses mRNA translation by stalling ribosomal translocation during elongation (By similarity). Reports are contradictory with regards to its ability to mediate translation inhibition of MBP mRNA in oligodendrocytes (PubMed:23891804). Also involved in the recruitment of the RNA helicase MOV10 to a subset of mRNAs and hence regulates microRNA (miRNA)-mediated translational repression by AGO2 (PubMed:14703574, PubMed:17057366, PubMed:25464849). Facilitates the assembly of miRNAs on specific target mRNAs (PubMed:17057366). Plays also a role as an activator of mRNA translation of a subset of dendritic mRNAs at synapses (PubMed:19097999, PubMed:19166269). In response to mGluR stimulation, FMR1-target mRNAs are rapidly derepressed, allowing for local translation at synapses (By similarity). Binds to a large subset of dendritic mRNAs that encode a myriad of proteins involved in pre- and postsynaptic functions (PubMed:7692601, PubMed:11719189, PubMed:11157796, PubMed:12594214, PubMed:17417632, PubMed:23235829, PubMed:24448548). Binds to 5'-ACU[GU]-3' and/or 5'-[AU]GGA-3' RNA consensus sequences within mRNA targets, mainly at coding sequence (CDS) and 3'-untranslated region (UTR) and less frequently at 5'-UTR (PubMed:23235829). Binds to intramolecular G-quadruplex structures in the 5'- or 3'-UTRs of mRNA targets (PubMed:11719189, PubMed:18579868, PubMed:25464849, PubMed:25692235). Binds to G-quadruplex structures in the 3'-UTR of its own mRNA (PubMed:7692601, PubMed:11532944, PubMed:12594214, PubMed:15282548, PubMed:18653529). Binds also to RNA ligands harboring a kissing complex (kc) structure,this binding may mediate the association of FMR1 with polyribosomes (PubMed:15805463). Binds mRNAs containing U-rich target sequences (PubMed:12927206). Binds to a triple stem-loop RNA structure, called Sod1 stem loop interacting with FMRP (SoSLIP), in the 5'-UTR region of superoxide dismutase SOD1 mRNA (PubMed:19166269). Binds to the dendritic, small non-coding brain cytoplasmic RNA 1 (BC1),which may increase the association of the CYFIP1-EIF4E-FMR1 complex to FMR1 target mRNAs at synapses (By similarity). Associates with export factor NXF1 mRNA-containing ribonucleoprotein particles (mRNPs) in a NXF2-dependent manner (By similarity). Binds to a subset of miRNAs in the brain (PubMed:14703574, PubMed:17057366). May associate with nascent transcripts in a nuclear protein NXF1-dependent manner (PubMed:18936162). In vitro, binds to RNA homomer,preferentially on poly(G) and to a lesser extent on poly(U), but not on poly(A) or poly(C) (PubMed:7688265, PubMed:7781595, PubMed:12950170, PubMed:15381419, PubMed:8156595). Moreover, plays a role in the modulation of the sodium-activated potassium channel KCNT1 gating activity (PubMed:20512134). Negatively regulates the voltage-dependent calcium channel current density in soma and presynaptic terminals of dorsal root ganglion (DRG) neurons, and hence regulates synaptic vesicle exocytosis (By similarity). Modulates the voltage-dependent calcium channel CACNA1B expression at the plasma membrane by targeting the channels for proteosomal degradation (By similarity). Plays a role in regulation of MAP1B-dependent microtubule dynamics during neuronal development (By similarity). Recently, has been shown to play a translation-independent role in the modulation of presynaptic action potential (AP) duration and neurotransmitter release via large-conductance calcium-activated potassium (BK) channels in hippocampal and cortical excitatory neurons (PubMed:25561520). Finally, FMR1 may be involved in the control of DNA damage response (DDR) mechanisms through the regulation of ATR-dependent signaling pathways such as histone H2AX/H2A.x and BRCA1 phosphorylations (PubMed:24813610).., [Isoform 10]: Binds to RNA homomer,preferentially on poly(G) and to a lesser extent on poly(U), but not on poly(A) or poly(C) (PubMed:24204304). May bind to RNA in Cajal bodies (PubMed:24204304).., [Isoform 6]: Binds to RNA homomer,preferentially on poly(G) and to a lesser extent on poly(U), but not on poly(A) or poly(C) (PubMed:24204304). May bind to RNA in Cajal bodies (PubMed:24204304).., (Microbial infection) Acts as a positive regulator of influenza A virus (IAV) replication. Required for the assembly and nuclear export of the viral ribonucleoprotein (vRNP) components..
Tissue specificity :Expressed in the brain, cerebellum and testis (PubMed:8401578). Also expressed in epithelial tissues (PubMed:8401578). Expressed in mature oligodendrocytes (OLGs) (PubMed:23891804). Expressed in fibroblast (PubMed:24204304). Expressed in neurons, Purkinje cells and spermatogonias (at protein level) (PubMed:8401578). Expressed in brain, testis and placenta (PubMed:8504300). Expressed in neurons and lymphocytes (PubMed:8504300)..
Subcellular locationi :Nucleus. Nucleus, nucleolus. Chromosome, centromere. Chromosome. Cytoplasm. Cytoplasm, perinuclear region. Cytoplasm, Cytoplasmic ribonucleoprotein granule. Perikaryon. Cell projection, neuron projection. Cell projection, axon. Cell projection, dendrite. Cell projection, dendritic spine. Cell junction, synapse, synaptosome. Cell projection, growth cone. Cell projection, filopodium tip. Cell junction, synapse. Cell junction, synapse, postsynaptic cell membrane. Cell junction, synapse, presynaptic cell membrane. Cell membrane. Cytoplasm, Stress granule.
IMPORTANT: For western blots, incubate membrane with diluted primary antibody in 1% w/v BSA, 1X TBST at 4°C overnight.